|

I'm a Senior Research Scientist at Relation, working on generative models of biological data. I am particularly interested in diffusion/flow models and ways to achieve generalisation and multimodality. Previously, I was at Valence Labs / Recursion, working on generative models for cell microscopy. I completed my PhD jointly at the University of Edinburgh (with C. Williams) and the Max Planck Institute for Intelligent Systems (with B. Schölkopf), working on generalisation and representation learning. During my PhD, I spent time at Google DeepMind, Meta AI (FAIR) and Spotify.
|

|
|
|
|
|
C Jones, E Noutahi, J Hartford*, C Eastwood* Preprint 2026 | Code We develop a simple and scalable recipe for training flow-matching models on cell-microscopy images. |
|
|
M Tschannen, C Eastwood, F Mentzer ECCV 2024 | Code We introduce generative transformers for real-valued vector sequences, rather than discrete tokens from a finite vocabulary. |
|
|
C Eastwood*, S Singh*, A Nicolicioiu, M Vlastelica, J von Kügelgen, B Schölkopf NeurIPS 2023 | Code We show how weak-but-robust predictions can be used to harness complementary spurious features in a new domain, boosting out-of-distribution performance. (Previously @ ICML 2023 Spurious Correlations Workshop) |
|
|
C Eastwood*, A Robey*, S Singh, J von Kügelgen, H Hassani, G J Pappas, B Schölkopf We learn predictors that generalise with a desired probability and argue for better evaluation protocols in domain generalisation. |
|
|
|
Z Ji, C Eastwood, A Goldenberg, PP Liang, J Hartford, RG Krishnan, E Noutahi Medical Imaging with Deep Learning 2026 We propose a contrastive multimodal method that only requires weak treatment-group labels rather than paired data. |
|
|
F Wenkel, W Tu, C Masschelein, H Shirzad, L Hodgson, I Bendidi, C Eastwood, ST Whitfield, C Russell, Y El Mesbahi, J Ding, MM Fay, B Earnshaw, E Noutahi, AK Denton Nature Biotechnology 2026 | Code We explore how knowledge graphs of gene-gene relationships can improve out-of-distribution predictions. |
|
|
K Donhauser, K Ulicna, GE Moran, A Ravuri, K Kenyon-Dean, C Eastwood, J Hartford ICML 2025 We extract interpretable biological concepts from microscopy foundation models. (Previously Oral @ NeurIPS 2024 workshop on Interpretable AI) |
|
|
C Eastwood, J von Kügelgen, L Ericsson, D Bouchacourt, P Vincent, B Schölkopf, M Ibrahim Preprint 2023 | Code We use data augmentations to disentangle rather than discard. |
|
|
C Eastwood*, A Nicolicioiu*, J von Kügelgen*, A Kekić, F Träuble, A Dittadi, B Schölkopf ICLR 2023 | Code We extend the DCI framework by quantifying representation "explicitness". (Previously @ UAI 2022 Causal Repr. Learning Workshop) |
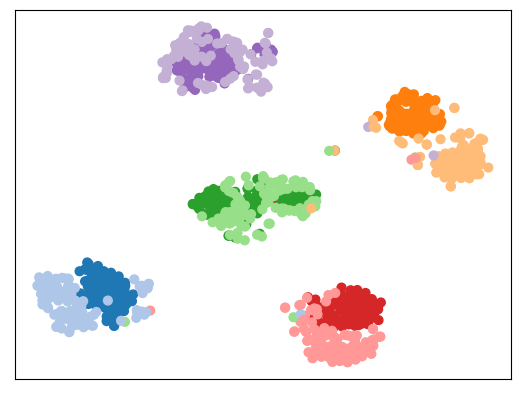  |
C Eastwood*, I Mason*, CKI Williams, B Schölkopf ICLR 2022 (Spotlight) | Code We address "measurement shift" (e.g., a new hospital scanner) by restoring the same features rather than learning new ones. |
|
|
C Eastwood*, I Mason*, CKI Williams NeurIPS 2021 ICBINB Workshop (Spotlight & Didactic Award) | Code | Video We use unit-level activation "surprise" to determine which parameters to adapt for a given distribution shift. |
|
|
N Li, C Eastwood, B Fisher NeurIPS 2020 (Spotlight) | Code | Video We learn accurate, object-centric representations of 3D scenes by aggregating information from multiple 2D views/observations. |
|
|
C Eastwood, CKI Williams ICLR 2018 | Code We propose the DCI framework for evaluating "disentangled" representations. (Previously Spotlight @ NeurIPS 2017 Disentanglement workshop) |